A template for reproducible papers
At the PINGA lab, we have been experimenting with ways to increase the reproducibility of our research by publishing the git repositories that accompany our papers. You can find them on our GitHub organzation. I’ve synthesized the experience of the last 4 years into a template in the pinga-lab/paper-template repository.
The template reflects the tools we’ve been using and the type of research that we do:
- Most papers are proposing a new methodology rather than the analysis of a dataset.
- There is always an application to a dataset to show the method works. We can’t always publish the data but we include it in the repository whenever we can.
- All papers include an implementation of the proposed method.
- Our code is usually written in Python and executed in Jupyter notebooks.
- The focus of the paper is usually on the methodology, not the code. As such, the code is more of a proof-of-concept than a full blown application or library.
- The paper itself is written in LaTeX with the source usually included in the repository.
This certainly won’t fit everyone’s needs but I hope that you can at least use
a few bits and pieces for inspiration.
Of course, the template code is open-source (BSD license) and you are free to
reuse it however you like.
The template includes a sample application to climate change data, complete
with a Python package, automated tests, an analysis notebook, a notebook that
generates the paper figure, raw data, and a LaTeX text.
Everything, from compilation to building the final PDF, can be done with a
single make command.

We’ve been using different versions of this template for a few years and I’ve been tweaking it to address some of the difficulties we encountered along the way.
- Running experiments in Jupyter notebooks can get messy when people aren’t diligent about the execution order. It can be hard to remember to “Reset and run all” before using the results.
- The execution was done manually so you had to remember and document in what order the notebooks need to be run.
- Experimental parameters (e.g., number of data points, inversion parameters, model configuration) were copied into the text manually. This sometimes led to values getting out of sync between the notebooks and paper.
- We only had integration tests implemented in notebooks. More often than not, the checks were visual and not automated. I think a big reason for this is the lack of experience in writing tests within the group and setting up all of the testing infrastructure (mainly how to use pytest and what kind of test to perform).
The latest update addresses all of these pain points. The main features of the new template are:
- Uses
Makefiles to automate the workflow. You can build and test the software, generate results and figures, and compile the PDF with a singlemakecommand. - A
Makefilefor building the manuscript PDF with extra rules for running proselint, counting words, and opening the PDF. - A starter conda environment for managing dependencies and making sure everyone gets the same version of the dependencies.
- Boilerplate instructions for downloading the code and reproducing the results.
- A
Makefilefor building the Python package, testing it with pytest, running static code checks (flake8 and pylint), and generating results and figures from the notebooks. - The code
Makefilecan run the notebooks usingjupyter nbconvertto guarantee that the notebooks are executed in sequential order (top to bottom). I would love to use nbflow but the SCons requirement puts me off a bit.makeworks fine and the basic syntax is easier to understand. - An example of using code to write experimental parameters in a
.texfile. The file defines new variables that are used in the main text. This guarantees that the values cited in the text are the ones that you actually used to produce the results.
This last feature is my favorite. For example, the notebook
code/notebooks/estimate-hawaii-trend.ipynb has the following code:
tex = r"""
% Generated by code/notebooks/estimate-hawaii-trend.ipynb
\newcommand{{\HawaiiLinearCoef}}{{{linear:.3f} C}}
\newcommand{{\HawaiiAngularCoef}}{{{angular:.3f} C/year}}
""".format(linear=trend.linear_coef, angular=trend.angular_coef)
with open('../../manuscript/hawaii_trend.tex', 'w') as f:
f.write(tex)
It defines the LaTeX commands \HawaiiLinearCoef and \HawaiiAngularCoef that
can be used in the paper to insert the values estimated by the Python code.
The commands are saved to a .tex file that can be included in the main
manuscript.tex.
Since this file is generated by the code, the values are guaranteed to be
up-to-date.
If you want to use the template to start a new project:
-
Create a new git repository:
mkdir mypaper cd mypaper git init -
Pull in the template code:
git pull https://github.com/pinga-lab/paper-template.git master -
Create a new repository on GitHub.
-
Push the template code to GitHub:
git remote add origin https://github.com/USER/REPOSITORY.git git push -u origin master -
Follow the instruction in the
README.md.
Alternatively, you can use the “Import repository” option on GitHub.
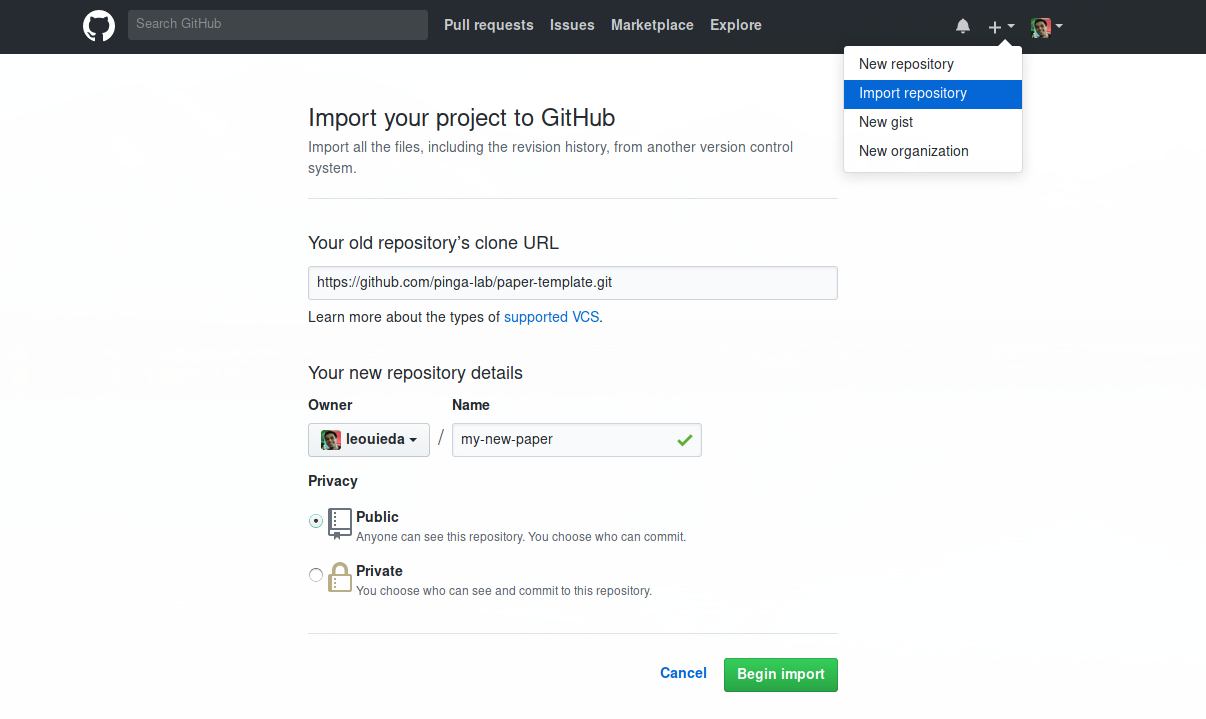
I hope that this template will be useful to people outside of our lab. There is definitely still room for improvement and I’m looking forward to trying it out on my next project.
What other features would you like to see in the template? I’d love to know about your experiences and workflows for computational papers.